|
9/3/2023 0 Comments Jmol viewer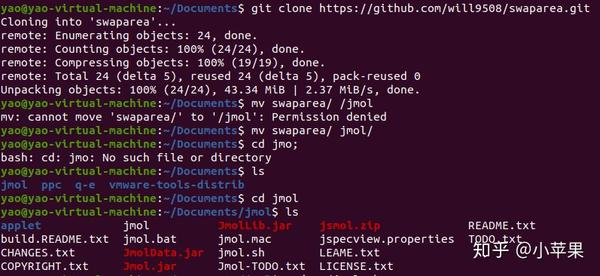
View the animated tutorial on the Protein Data Bank website. The website has an animated tutorial that introduces the viewer to some the features of the new interface for the Protein Data Bank. The Protein Data Bank website currently contains close to 40, 000 structure files along with a wealth of information on structural biology and bioinformatics. 
The RCSB is a collaboration between Rutgers University, the University of California-San Diego and the University of Wisconsin-Madison. The best source of structure files for biological macromolecules is the Protein Data Bank, which is a Federally supported database that is hosted by Rutger University and is managed by the Research Collabratory for Structural Bioinformatics (RCSB). We will now obtain a structure file that we can work with with Jmol. Placing the structure files in the same folder as the "Jmol.jar" application will make them easier to find and open.Ĭreate a folder on your "username_X" share and place a copy of the "Jmol.jar" in the folder. To run Jmol as a standalone application, the only file you need is "Jmol.jar":įor this tutorial, we will copy the "Jmol.jar" application file to a separate folder in which we will also place the structure files for the molecules that we wish to look at with Jmol. There are a number of files in the the folder, most of which are used to serve webpages containing Jmol applets. the version that will be used in this tutorial is "Jmol-10.3.1". Go to the Jmol website and download a copy of the latest version of Jmol.Īfter download the download file and extracting it you should endup with a folder called "Jmol-x.x.x", where x.x.x is the version number. Before you can get started with learning how use Jmol, you must first obtain a copy of the program, along with a file containing the atomic coordinates of a molecule that wish to view with Jmol.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
